Hardware Solutions
Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Immunology
Researchers led by Dr. Maria Kaparakis-Liaskos from Monash Institute of Medical Research in Australia are studying the nucleotide-binding oligomerization domain-1 (NOD1) receptor, which senses the presence of bacteria and then stimulates an immune response. To gain a better understanding of exactly how and where NOD1 initiates an immune response, the researchers used Imaris to visualize the location and interactions of NOD1 with its ligand inside a cell.
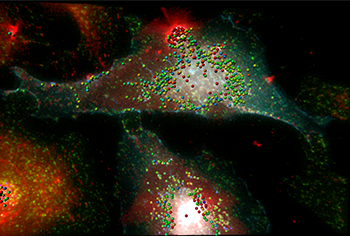
The 3D fluorescence image shows HeLa cells in the background expressing NOD1-CFP (cyan) stimulated with peptidoglycan-containing Helicobacter pylori outer membrane vesicles (red). A co-localization color-coded map of NOD1 (cyan) at early EEA1-labeled endosomes (green) with outer membrane vesicles (red) in HeLa cells is shown. Co-localization is indicated by the spheres, where red indicates relatively high levels of co-localization in contrast with locations of lesser levels of co-localization shown in cooler colors. Image by Camden Lo, Monash Micro Imaging.
NOD1 is an important part of our innate immune system. This pattern recognition receptor senses the presence of Gram-negative bacteria by detecting peptidoglycan, which forms a mesh-like layer on the outside of bacteria. Studies have identified NOD1 is involved in a broad array of biological functions.
“Despite the identification of NOD1 as being the receptor for bacterial peptidoglycan in 2003, the interaction between NOD1 and peptidoglycan has not been previously visualized,” said Kaparakis-Liaskos. “Expanding on our limited understanding of the intracellular location of NOD1 and its interaction with peptidoglycan will enable the design and development of NOD1 antagonists that have the potential to prevent NOD1-driven inflammatory and autoimmune disorders in humans.”
Power and flexibility
Kaparakis-Liaskos explained that Imaris provided the power and flexibility that they needed to analyze and quantify colocalization of the fluorescently labeled molecules under study. The investigators used Imaris software at the Monash Micro Imaging (MMI) core facility, which specializes in microscopy and image analysis as part of the Monash Institute of Medical Research and Monash University. MMI is a member of Bitplane’s network of Advanced Imaging Centres, which help shape Bitplane’s software development by providing information on key trends and needs.
“We have a very constructive relationship with Bitplane in terms of technical assistance, feedback, and capabilities development,” said Dr. Camden Lo, who was part of the research team and also an MMI imaging research fellow. Drs. Aaron Irving and Lo were responsible for performing all of the Bitplane analysis of the images for the study.
To visualize NOD1, the researchers used outer membrane vesicles from pathogenic bacteria to deliver peptidoglycan into the host cell cytosol. They collected three-dimensional fluorescence image stacks at 0.2-micron intervals and used Imaris to model the cellular compartments with the Surfaces and Cell analytical modules. They also used the built-in Imaris co-localization – Coloc to assess for the potential of peptidoglycan and NOD1 interactions. With this approach, they masked for NOD1-containing vesicles and then calculated the percentage that co-localized with early endosome signals. Early endosomes are a type of membrane-bound endocytic organelle that serves as a focal point of the endocytic pathway.
“We observed NOD1, its ligand peptidoglycan, and the NOD1 adaptor protein RIP2 all co-localizing at early endosomes, said Kaparakis-Liaskos. “This is the first time the receptor NOD1 has been seen interacting with its ligand and RIP2 within a defined intracellular location.”
Overall the findings demonstrate that NOD1 detects peptidoglycan within early endosomes, thereby promoting RIP2-dependent degradation of bacteria and inflammatory signaling in response to bacterial infection.
Author: Maria Kaparakis-Liaskos and colleagues, Monash Institute of Medical Research
Category: Case Study
