Hardware Solutions
Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Neuroscience
There is mounting evidence that protein synthesis and its regulation play a pivotal role in numerous neural processes such as the formation of new synaptic contacts and long-lasting forms of synaptic plasticity. These processes, which are necessary for learning and memory, involve changes in the cellular proteome via protein synthesis, its posttranslational modifications and ultimately degradation.
Researchers led by Dr. Daniela C. Dieterich and Erin M. Schuman of the California Institute of Technology and Leibniz Institute for Neurobiology as well as members from Institut National de la Santé et de la Recherche Médicale (INSERM) have developed a technique called fluorescent non-canonical amino acid tagging (FUNCAT) that allows visualization of changes in protein synthesis in the proteome on the time scale of minutes.
“We know that some synaptic proteins can be synthesized only in the soma whereas some synaptic proteins appear to have a local source, but we do not understand the factors governing either the location of synthesis of synaptic proteins or their local fate, and to which quantities these proteins are synthesized,” she said.
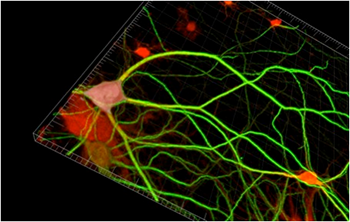
FUNCAT metabolically labels newly synthesized proteins by incorporating non-canonical amino acids into the proteins and then detecting them with chemoselective fluorescent tags using click chemistry. “FUNCAT enables the visualization of newly synthesized proteins in a highly specific fashion utilizing the cell’s own translation machinery and artificial amino acids bearing a functional azide moiety. Applying the artificial amino acid directly on a dendrite or on a soma can monitor local changes in dendritic or somatic, respectively, protein synthesis,” Dr. Dieterich said.
The researchers synthesized the fluorescent tags Texas Red-PEO2-alkyne (TRA) and 5'-carboxyfluorescein–PEO8–azide (FLA), which would bind to the modified amino acids azidohomoalanine (AHA) homopropargylglycine (HPG) that were incorporated into proteins. They then used the technique to visualize newly synthesized proteins in dissociated primary hippocampal neurons, acquiring images with a confocal laser scanning microscope and performing postacquisition processing and analysis with ImageJ and Imaris software. Imaris let the researchers work with masks, which allowed them to distinguish between neuronal and glial cell types and to analyze the signal intensities for newly synthesized proteins separately in cell somata and dendrites in three dimensions, said Dr. Dieterich.
To study the diffusion properties of newly synthesized proteins the researchers analyzed unfiltered and unprocessed stacks of straightened dendrites with Imaris. They calculated the TRA signal intensities per volume by creating a three-dimensional mask using corresponding stacks of straightened dendrites that had been immunostained for the dendritic marker protein MAP2. They then measured the MAP2-defined volume and the unprocessed TRA signal intensities across all the stacks that were included by the MAP2-defined mask. The researchers were able to detect newly synthesized proteins in neuronal somata after 10 minutes of exposure to AHA or HPG. Longer exposure increased the intensity of the fluorescent signal in cell bodies and fluorescence could be seen in the dendrites.
With FUNCAT the researchers were able to detect changes in protein synthesis, analyze the fate of proteins, and visualize the dynamics of newly synthesized membrane proteins with single particle tracking and quantum dots. They say that this method could be adapted for fluorophores used with STED high-resolution microscopy, and they foresee FUNCAT being used to directly observe protein synthesis in neural circuits involved in learning and memory.
Research Paper: In situ visualization and dynamics of newly synthesized proteins in rat hippocampal neurons, Nature Neuroscience, 13, 897–905 (2010), doi:10.1038/nn.2580.
Author: Dr. Daniela C. Dieterich, Erin M. Schuman and colleague, California Institute of Technology and Leib
Category: Case Study
