Hardware Solutions
Applications
 Part of the Oxford Instruments Group
Part of the Oxford Instruments Group
Embryology
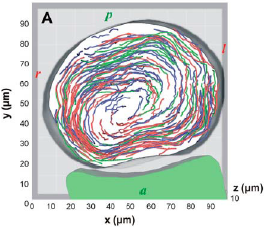
Living systems rely on fluid dynamics from embryonic development to adulthood. One well known example is the specification of the left-right axis in vertebrates, where beating cilia generate a directional flow necessary for breaking the embryonic symmetry in the so-called left-right organizer.
To visualize biological fluid flow, devising the proper labelling method compatible with both normal biology and in vivo imaging remains a major experimental challenge. In the group of Professor Scott Fraser, Dr Willy Supatto and his colleague Dr Julien Vermot used an all-optical procedure combining femtosecond laser ablation, fast confocal microscopy and 3D-particle tracking to label, image and quantify cilia-driven flow generated within the Kupffer’s vesicle (KV), the left-right organizer of zebrafish embryos.
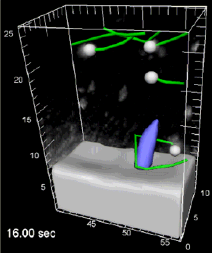
The group used Imaris to quantify this fluid flow generated within the KV. After seeding the fluid with fluorescent particles, the 3D-particle motion over time was tracked automatically with Imaris and corrected manually. ImarisXT was used to export the data in Matlab and fully characterize the flow with further analysis in Matlab. The present study show that the flow driven by beating cilia is breaking the left-right symmetry and provides an experimental validation of flow features predicted from simulations.
Movie 1 shows a dynamic version of Figure 1, exhibiting the track occurrence and the 3D particle motion over time in the vesicle. Movie 2 shows a dynamic version of Figure 2, showing the movement of particlules trapped in the vortical flow surrounding a cilium.
This study is the first quantifying the 3D flow surrounding a beating cilium inside a living organism and provides new experimental access to questions of microscopic fluid dynamics in vivo. Such detailed descriptions of cilia-driven fluid movement will be valuable in unraveling the relationships between flow and signal transduction crucial for maintaining the asymmetry of the embryo. In general, this approach opens new opportunities for investigating microscopic flow in living tissues.
Author: Dr. Willy Supatto and colleagues, Biological Imaging Center California Institute of Technology
Category: Case Study
